Silencing genes — to understand them
How scientists block cells’ genetic messages — and why
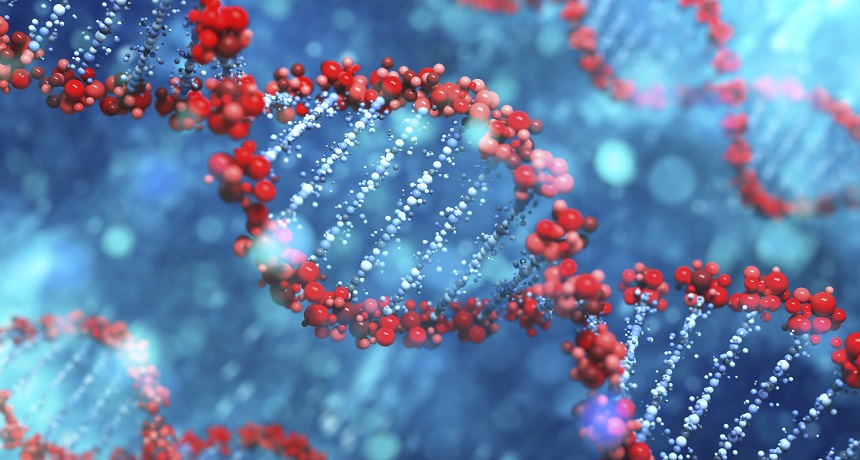

Scientists have identified all of the genes that lie within our DNA, depicted here. Now they’re probing what all of those genes do — and often using RNAi to find out.
SilverV/ iStockphoto
The DNA in each of us hosts about 21,000 genes. Their blueprints are coded in the roughly 3 billion rungs of our DNA’s ladder-like structure. The Human Genome Project finished decoding all of those genes in 2003. That task took hundreds of scientists more than 12 years.
Scientists are now working fast and furiously to learn what each identified gene does. Their answers will help science better understand how cells work. The knowledge also could help doctors better treat disease.
“One great way to learn what cells do normally with all their different genes is to turn off those genes one at a time,” explains Craig Mello. He’s a geneticist at the University of Massachusetts Medical School in Worcester.
David Root is a geneticist at the Broad Institute in Cambridge, Mass. To show how turning off genes can help, Root likens our complex genome — the complete encyclopedia of our genes — to the main electrical box in your house, full of fuses or circuit breakers. If none of the circuits feeding the box was labeled, he says, “You might turn each [circuit] off and see which of your lights go out or which of your appliances turn off.”
Scientists can trigger RNAi too. And by doing so, “We can trick the cell into shutting down any one gene — pretty much any gene in the whole genome — any time we want,” says Mello.
Turning off genes this way allows scientists to find out what they do when they’re on. Read on to see how RNAi works and what new work that uses it shows.
Our genetic alphabet
Cells have two kinds of genetic material: DNA and RNA. Our genes reside on DNA. Its long molecules resemble twisted ladders.
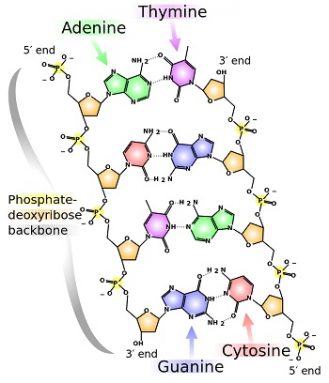
Each rung in DNA’s ladder-like structure is a pair of two chemicals — called nucleotides. Together they make up a base pair. The four nucleotides in DNA are known as A, T, C and G. They stand for adenine, thymine, cytosine and guanine. The nucleotide attached to each long side on the twisted ladder must pair with a particular one on the other side. All A’s, for instance, pair with T’s. C’s will only pair with G’s.
“DNA is a lot like the hardware in a computer. It’s something you’re born with,” says Mello. “Every time the cell reproduces,” he explains. “it reproduces a full set of these instructions that allow the cell to do its thing.”
Putting DNA’s instructions to work is one job of RNA.
“RNA is also a string of four different units, and it matches up one-to-one with DNA,” Root says. Three of those units are the A, C and G found in DNA. But in RNA, “instead of the T, you’ve got a U.” That U stands for uracil. Like T, it always — and only — binds to A.
While some types of RNA are double-stranded, most forms have a single strand of nucleotides. Different types of RNA do different jobs.
Messenger RNA, or mRNA, carries a gene’s instructions to structures in a cell known as ribosomes (RY-boh-sohmz). This information tells other RNA molecules in those ribosomes to make some particular protein. Those proteins will carry out most of a cell’s functions.
But not every cell needs to do every task. For example, nerve cells and skin cells have very different tasks. Nor must each cell work at the same time.
Instead, each cell expresses — or turns on — the genes that it needs for a particular job at some particular time. Different RNA molecules help control that process.
One way RNA does this is with proteins called Argonautes. These specialized proteins, named for a band of heroes in Greek mythology, bind to small RNA molecules. They help coordinate other proteins in events that result in the eventual silencing of a gene.
“Argonautes are very much like the search engines we use on our Web browser,” says Mello. The small RNAs function like the query for a search. They resemble the few words that you might type into a search window to find detailed information about a topic. Argonautes work with bigger chunks of information to find matches for the search query. And, Mello notes, “Argonautes can do this very rapidly.”
When the short piece of RNA bound to the Argonaute matches up with part of a specific messenger RNA, the Argonaute protein cuts the mRNA. The mRNA falls apart, and that destroys its message. Thus, the ribosome never gets the instruction to make a certain protein.
This process is RNA interference, or RNAi. It keeps cells from making proteins that aren’t needed at that time. Cells also use RNAi “to clean up any RNA that shouldn’t be there or that isn’t perfect,” explains Lisa Stanek. She’s a neuroscientist at Genzyme Corp. in Cambridge, Mass.
The process happens naturally in animals, plants, fungi and even some bacteria. Says Mello: “We think RNA interference is at least a billion years old!”
Hijacking the process
Since the first news about its discovery in 1998, researchers have wanted to use RNAi to study cells. To do that, scientists learned how to “hijack it and reprogram it,” says Christof Fellmann. He’s a molecular biologist at Mirimus Inc., in Cold Spring Harbor, N.Y.
Scientists can trigger RNAi, for example, by introducing synthetic double-stranded RNA molecules. Their shape “resembles a short hairpin,” Fellmann explains.
“Nature had millions of years of evolution to fine-tune these things,” Fellmann notes. Making artificial triggers for RNAi, he believes, could help in both research and medicine.
Of course, triggers need a way into cells. One trick: Stuff them into a virus and then let the germ ferry the molecule into target cells.
“Viruses’ jobs are to infect cells,” explains Stanek at Genzyme. First, bioengineers take out all genes that might make someone sick. Then they tinker with the germ. “We engineer the virus to deliver our RNAi,” she says.
Triggering RNAi to occur on demand
But that may not always work. Some immune cells destroy viruses, for instance. So researchers working with the Broad Institute found a way to insert the triggers with nanowires. They injected teensy wires, like needles, into cells.
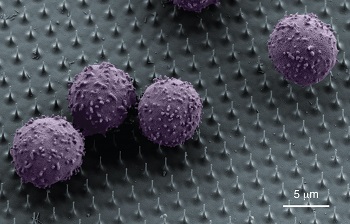
To bind to its targets, an RNAi trigger needs a negative ionic charge, he explains. (An ion is an atom or molecule with an electric charge due to the loss or gain of one or more electrons. Something with a negative charge has extra electrons.) But immune responses in humans often keep negatively charged molecules out of cells.
To avoid that, Dowdy and his team neutralized the charge of their RNAi trigger. They designed molecules with an added chemical group, called a phosphotriester (FOSS-foe-try-ESS-tur). This acts like a mask so that the RNAi trigger molecule can sneak into cells. Once inside, a cellular enzyme strips away the added chemical group.
“This little Pac-Man enzyme sees it right away, and it starts chewing off this phosphotriester group,” says Dowdy. The remaining molecule has a negative charge, so it can now trigger RNAi. Dowdy’s team reported this technique in the November 2014 Nature Biotechnology.
Getting an RNAi trigger into a cell is only part of the task, though. “To figure out what a gene can do, you actually have to do that experiment over and over again, because the answer turns out to be different in different kinds of cells and conditions,” explains Root at the Broad Institute.
“You don’t necessarily learn about what a gene does in neurons by changing that gene in an immune cell,” he notes. You can’t tell, “even by looking at a neuron in isolation,” he adds. And changing the environment that surrounds a cell may change the way it behaves when a gene is silenced.
Fortunately, automated equipment lets researchers test many kinds of cells and triggers at once. Tests don’t need much space, either.
“On your palm you can fit several hundred RNAi experiments,” says Anastasia Eskova. She’s a cell biologist at the Max Planck Institute for Developmental Biology in Tübingen, Germany.
Exploring cells, treating disease and more
RNAi helped show Eskova and her colleagues how some cells move. Those cells form — and then later eliminate — tiny anchors on their outer covering, called a membrane. Then they absorb the anchors at one end before making new anchors at the other end. This allows them to move like little caterpillars. Cancer cells do this, for instance, as the disease spreads. Immune cells also do this to stalk harmful invaders.
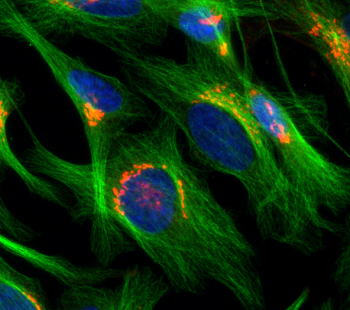
“We turned off, one by one, almost 400 different genes,” says Eskova. About 122 of them turned out to play some role in absorbing and moving the anchor material. One gene even turned out to control the transport of material across the cell for reuse. Scientists hadn’t known about that before. Her team published its findings in the June 2014 Journal of Cell Science.
Other RNAi research aims to treat disease. Genes often mutate, which means they change. Some of those changes may be harmful. They might slow or alter normal life functions. But RNAi could keep the mutant cells from making one or more proteins that would otherwise go on to cause trouble.
Stanek and others at Genzyme hope to help patients with a fatal genetic brain illness known as Huntington’s disease. Just one mutant copy of a single gene causes the disease. The mutation changes the gene’s code into the blueprint for making a protein that attacks the nervous system. Symptoms don’t tend to emerge until someone is 30 to 40 years old. Uncontrolled writhing and memory problems eventually develop. Patients become depressed and quickly upset.
Stanek’s team used RNAi in mice to curb expression of the gene in Huntington’s disease. Scientists still don’t know why patients might need the normal copy of that gene, though. So, Stanek’s team limited the amount of RNAi triggers so they wouldn’t shut it down entirely.

Other diseases often involve two or more genes. Still, RNAi might help. For example, Root and other researchers are looking for better gene targets to treat cancer.
One study found a protein on which certain leukemia cells depend. Normal blood cells don’t need the protein. Knocking out the gene for that protein might treat the disease, concludes Root and other scientists. They described their discovery in the July 2013 Cancer Cell.
Another study found genes that can switch off the killing action of certain cells of the immune system. Signals from cancer cells can sometimes trigger those off switches. Then the immune cells don’t attack the cancer. Knowing about that might lead to new treatments.
“You can try to block the switches, and not let the cancer use them,” Root explains. Then the immune cells should keep going and fight the cancer. Other research has used a similar approach for some new treatments, he notes. Root’s team shared its findings in the February 6, 2014 issue of Nature.
All in all, RNAi is a powerful tool. “We can now potentially fix certain types of genetic disease,” Mello says.
Beyond that, RNAi and other processes may help science understand how life works at the most basic level. And with so much left to learn, today’s young people may be the ones who ultimately make some of these important discoveries.
“Everything alive on this planet is so incredibly sophisticated,” Mello says. “It’s so cool.”
Word Find (click here to enlarge for printing)